-Search query
-Search result
Showing 1 - 50 of 733 items for (author: luo & t)

EMDB-45206: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45240: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45252: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45973: 
Bufavirus 1 at pH 2.6
Method: single particle / : Gulkis MC, McKenna R, Bennett AD

EMDB-45176: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45180: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c47: 
TRRAP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

PDB-9c4b: 
Second BAF53a of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
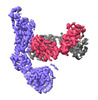
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
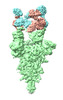
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
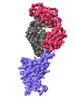
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
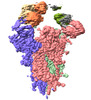
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
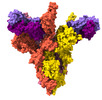
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-43374: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
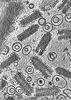
EMDB-43375: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
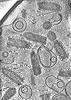
EMDB-43376: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L

EMDB-43377: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
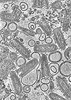
EMDB-43378: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
EMDB-43379: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
EMDB-43380: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L
EMDB-43381: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L

EMDB-43383: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L

EMDB-43384: 
CryoET of VSV incubated with liposomes at pH 5.5
Method: electron tomography / : Si Z, Xia X, Tang S, Milojevic L

EMDB-19822: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+bromosterol (DOPC, DOPE, DOPS, bromo-ergosterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18307: 
Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

EMDB-18308: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18309: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18310: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18311: 
Compact state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18312: 
Stretched state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qb7: 
Pil1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

PDB-8qb8: 
Lsp1 in native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

PDB-8qb9: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbb: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

PDB-8qbd: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
